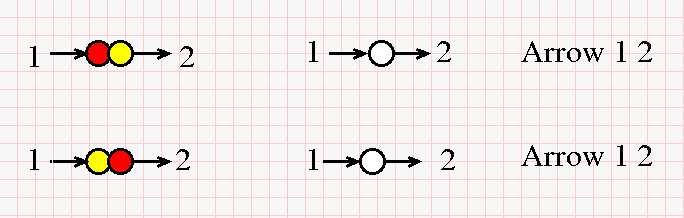Cite as: M. Buliga, Chemlambda strings, version 15.04.2017, http://imar.ro/~mbuliga/chemlambda_strains.html
Chemlambda strings is an alternative version of the chemlambda artificial chemistry which may be closer to real chemistry than the original. It does bring us closer to the dream of building a real molecular computer in the sense described in this article.
Assume we have a provision of asymmetric molecules, of three types, described as red, yellow and blue. They are asymmetric in the sense that we may identify a part which is called, conventionally, "in" and a part which is called "out", so that there is an unambiguous way to tell the "in" from the "out", as well as the possibility to identify a sense from "in" to "out". The color (red, yellow, blue) may be imagined as representing different chemical composition, or perhaps a part of the molecule which is different, in three ways.
These molecules can connect one with another in the following ways. As concern the red and yellow molecules, they can make long chains, or strings of alternating red and yellow molecules, made by connecting the "out" of a molecule with the "in" of the next one, in the sense defined by their asymmetry. A blue molecule can connect via the "out" part with the "in" part of a red or yellow molecule. Any other connection between "in" and "out" parts is prohibited.
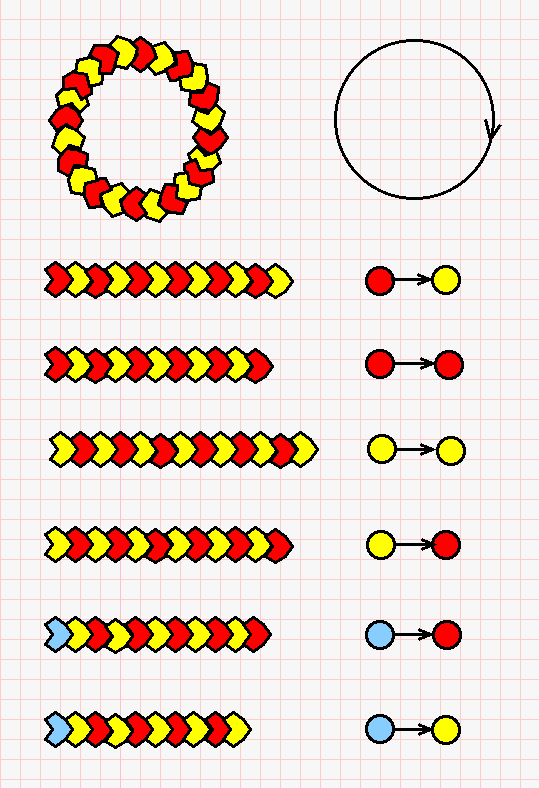
These strings of molecules can be described by indicating the color of the ends of the strings, if any. The possible strings are described in the next picture (click on it for a bigger version).
The red and yellow molecules have also a different part, let's call it "middle". Any two red or yellow molecules can connect via a "middle" - "in" or a "middle" - "out" connection, provided that the molecule which brings the "middle" part is not an end of a string. Any blue molecule can connect it's "in" part with a "middle" part of a red or yellow molecule which is not an end of a string.
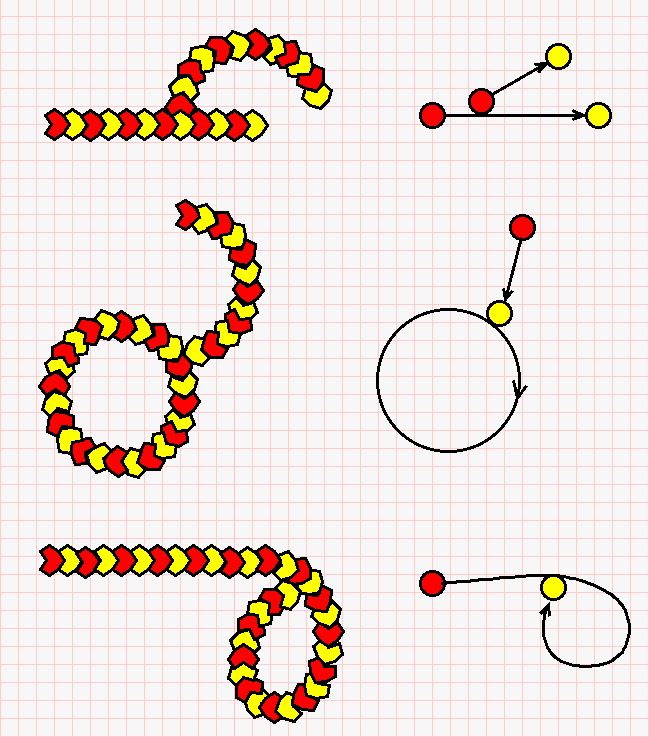
This gives more complex structures, made of one or several strings. We describe these connections as a tangency between an extremity of one string and (the bulk of) another string. Some examples are given in the next figure (click on it for a bigger version).
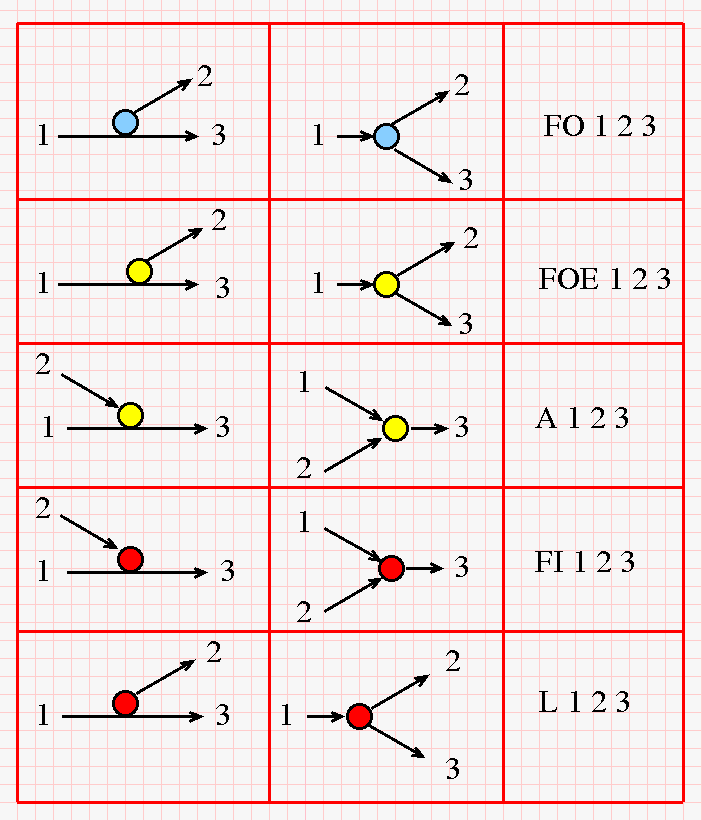
Each "middle"- * connection encodes a trivalent node of an oriented trivalent ribbon graph. More precisely, there is the following correspondence between the possible "middle"-* connections of strings and the nodes which compose chemlambda graphs. (Click on the figure for a bigger version.)
The figure has 3 columns. In the first column is a possible connection of strings, with the strings pieces around the connection numbered with "1", "2", "3". In the second column we see the corresponding chemlambda node. In the third column is the "mol" encoding of the node as used in the mol notation from chemlambda. With respect to the color codes used in chemlambda, here we have a slightly different convention, namely: (first line) a fanout "FO" node is colored with blue, instead of green in the original version, an application "A" node is colored with yellow, instead of green in the original version.
As for the auxiliary "Arrow" 2-valent nodes which are used in the original chemlambda, they are trivial here, because of the way a string is built. Therefore there will be no "Arrows" in this version, nor the corresponding COMB rewrites of chemlambda. As explained in the next figure, an "Arrow" node just expresses the fact that a string can be seen as two strings connected end-to-end. (Click on the figure for a bigger version.)
The graph rewrites of the original chemlambda are seen as chemical reactions of the form:
(Pattern before the graph rewrite) + (rewrite enzyme) - - - > (Pattern after the graph rewrite) + (rewrite enzyme)
An interesting feature of the Chemlambda strings version described here is that we can renounce at these invisible rewrite enzymes. Moreover we can make the chemical reactions implementing the chemlambda rewrites to conserve the strings, equivalently to conserve the nodes and arrows.
This is very attractive from the point of view of real chemistry. In this version of chemlambda we replace the rewrite enzymes by strings too.
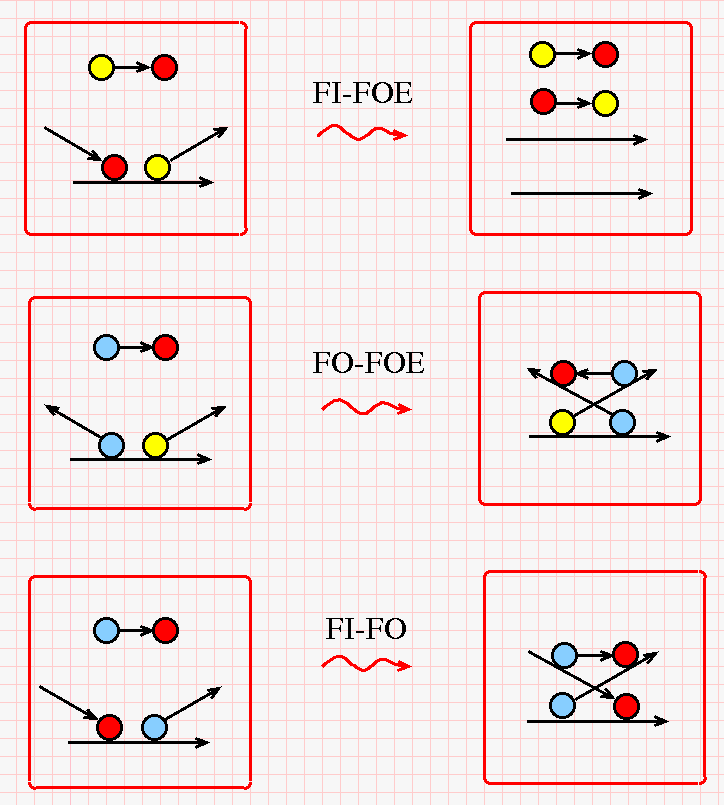
The most important rewrites of chemlambda involve the nodes "FI", "FO" and "FOE". In the next figure is given the chemlambda strings version, as a chemical reaction which transforms the left side box into the right side box. Each reaction has a name which corresponds with the name of a chemlambda graph rewrite. (Click on the figure for a bigger version.)
Somme comments are needed. The FI-FOE rewrite in the original chemlambda uses "Arrow" elements, which are then erased by the COMB rewrites. Here we see that a free yellow-red string acts indeed as an enzyme (because it is present in both sides of the reaction). Instead of "Arrows", the reaction result contains also a free red-yellow string, which will appear in other chemical reactions.
The FO-FOE and FI-FO rewrites are in the family of "distributive", or DIST rewrites of chemlambda. Here they appear as a free blue-red string which (1) permutes two neighboring string ends which sit on a string, then (2) the free blue-red string glues itself on the permuted strings.
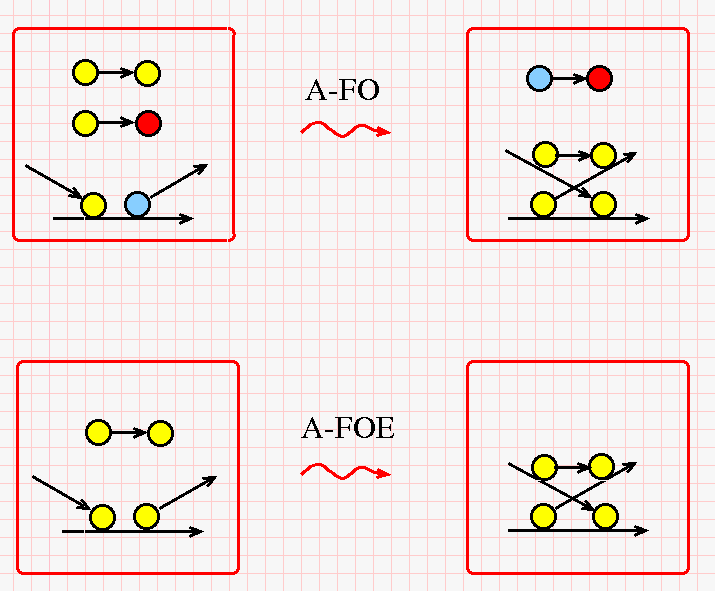
The DIST rewrites for the application "A" node in chemlambda are given in the next figure (click on it for a bigger version).
The A-FO rewrite needs a free yellow-yellow and a free yellow-red string and it produces a free blue-red string. Otherwise is the same permute-then-glue reaction. The A-FOE reaction needs only a free yellow-yellow string.
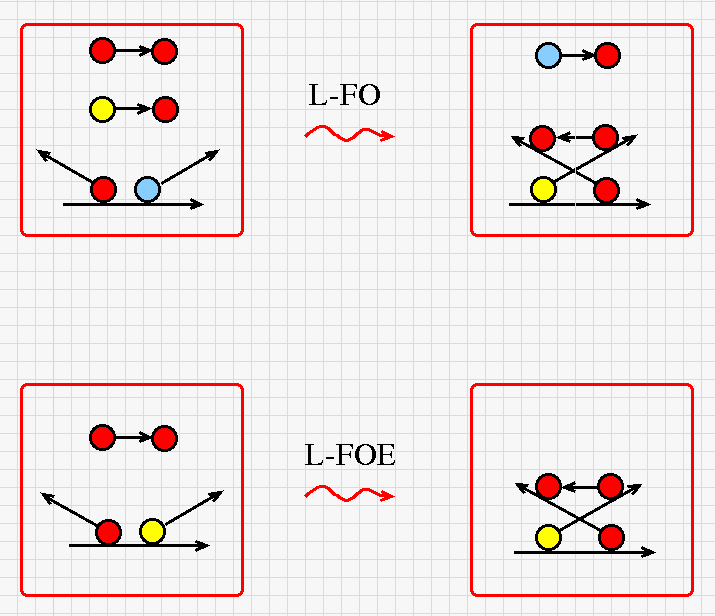
The DIST rewrites for the lambda "L" node in chemlambda have a similar pattern. This time red-red and yellow-red free strings are needed and a blue-red free string is produced. (Click on the figure for a bigger version.)
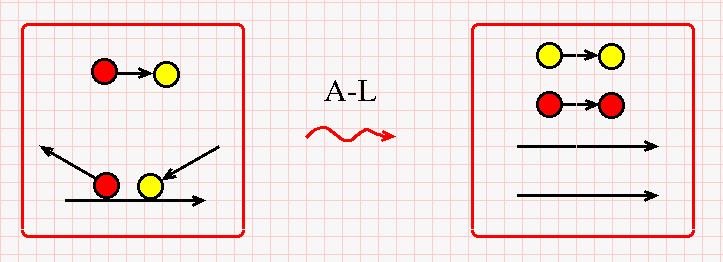
Finally, here is the A-L rewrite (inspired from the famous BETA rewrite from lambda calculus).
It is almost similar with the FI-FOE rewrite, but the A-L reaction needs a free yellow-red string and produces a free yellow-yellow string and another free red-red string. No "Arrow" nodes are needed.
In future versions we shall explore chemlamdba strains in relation with various phenomena in chemlambda, in particular chemlambda quines and kinesins.
[1] (github) - M. Buliga, Molecular computers (2015)
[2] (journal) - M. Buliga, Build a molecular computer. Journal of Brief Ideas, 2015.
[4] (MIT Press free download) (arxiv) - M. Buliga, L.H. Kauffman, Chemlambda, universality and self-multiplication (2014), Artificial Life 14, Proceedings of the Fourteenth International Conference on the Synthesis and Simulation of Living Systems
Edited by Hiroki Sayama, John Rieffel, Sebastian Risi, Rene Doursat and Hod Lipson, p. 490 - 497
[5] (github) - M. Buliga, The Chemlambda repository. For using the repository see the README of the active branch.
[6] (journal) (arxiv) - M. Buliga, Graphic lambda calculus. Complex Systems 22, 4 (2013), 311-360
[5] (github) - J. King, The Chemlambda-hask repository